Systems and Methods to Automatically Detect Ellipsoid Zone Loss in SD-OCT Imaging
Summary:
The National Eye Institute (NEI) seeks research co-development partners and/or licensees for an automatic deep learning-based algorithm to detect and quantitate ellipsoid zone (EZ) loss in Spectral Domain Optical Coherence Tomography (SD-OCT) images.
Description of Technology:
The present disclosure generally relates a method of automatically detecting ellipsoid zone (EZ) loss in spectral domain optical coherence tomography (SD-OCT) imaging. EZ band represents outer segments of photoreceptors in the retina, and its loss reflects a deterioration of the photoreceptors. EZ loss has been proposed to be evidence of progression of several retinal degenerative diseases including, but not limited to, retinitis pigmentosa and hydroxychloroquine (HCQ)-induced retinal toxicity. HCQ is a first-line drug used to treat autoimmune diseases such as systemic lupus erythematosus, Sjögren’s syndrome, and rheumatoid arthritis. One of the major side-effects that can occur in long-term users of HCQ, is EZ loss that can result in retinal toxicity and permanent damage to photoreceptors and retinal pigment epithelium (RPE), eventually leading to irreversible loss of central vision. The American Academy of Ophthalmology (AAO) recommends two main screening modalities including SD-OCT imaging and functional tests such as visual fields with the goal of recognizing early definitive signs of HCQ-induced retinal toxicity to prevent vision loss. Although this side-effect is estimated to occur in 7.5% of patients taking the drug for more than 10 years, we currently have no treatment for this serious side effect that tends to continue even after the cessation of the drug.
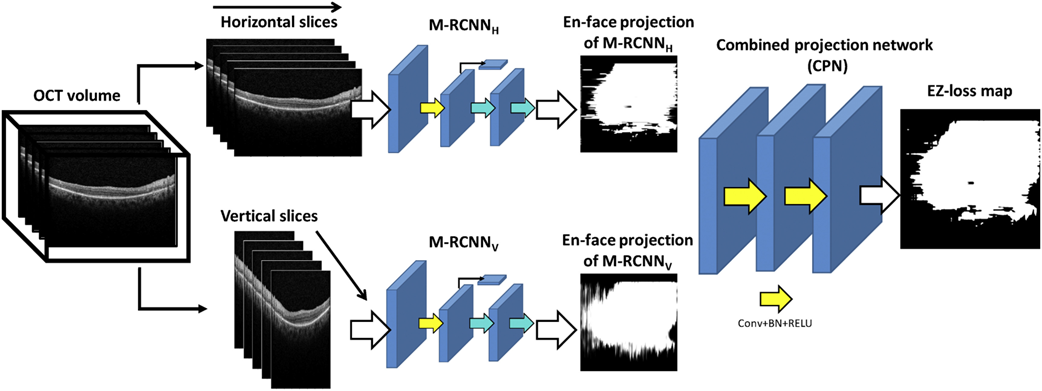
Researchers at the NEI have developed a method that can automatically detect and quantitate EZ loss in SD-OCT images immediately after image acquisition. The method includes a deep learning framework with a two-step approach. In the first stage, the method detects and annotates EZ loss regions in individual OCT B-scans. A 2D map is constructed twice in a dual architecture to enhance robustness, where horizontal and vertical slices extracted from the 3D image are trained separately. The second stage of the model operates on these two 2D maps and estimates the final EZ loss map representing the 3D OCT volume. Compared to other screening methods, the algorithm demonstrated excellent performance in diagnosing toxicity even as a stand-alone test, with an F1 score, a measure of test accuracy, of 0.91. This indicates the utility of the tool in assisting with screening for toxicity in an automatic, accurate, time-effective, cost-effective, and objective manner. Addition of this methodology onto current SD-OCT screening could assist the clinician in making diagnostic and treatment decisions immediately after SD-OCT acquisitions.
Protected claims for this invention include the method of detecting and outlining the region of EZ loss directly from acquired OCT images, associated algorithms, and the device containing these algorithms and capabilities. This technology is available for licensing.
Potential Commercial Applications:
- Methodology to integrate with SD-OCT imaging for screening applications for different retinal degeneration diseases
- Implementation of this automatic algorithm also in screening outside of ophthalmology offices (OCTs become more ubiquitous in internal medicine settings)
- Quantitative data produced from the algorithms could provide surrogate end points for use in clinical trials and interventional studies aimed at halting progression of degenerative changes
Competitive Advantages:
- Accurate and robust method: Provides excellent performance in diagnosing toxicity as a stand-alone test and compared to qualitative inspection of SD-OCT images
- Time-efficient (can evaluate SD-OCT volumes): Fully automatic and available within seconds after SD-OCT acquisition
- Objective: Performs as well as human grading in eyes that demonstrate even small amounts of EZ loss
- Does not require segmentation of intact layers, avoiding failures in cases of layer deterioration
- Does not require a large number of training examples, useful for rare diseases